GenomicRanges
What is GenomicRanges?
The GenomicRanges package serves as the foundation for representing genomic locations within the Bioconductor project. This R package lays the groundwork for genomic analysis by introducing three classes (GRanges, GPos, and GRangesList), which are used to represent genomic ranges, genomic positions, and groups of genomic ranges.
The human genome comprises roughly 3 billion base pairs organized linearly on 23 pairs of chromosomes. An intuitive way to represent our genome is to use a coordinate system: “chromosome id” and “position along chromosome”. An annotation like chr1:129-131 would represent the 129th to the 131st base pair on chromosome 1.
The ability to efficiently represent and manipulate genomic annotations and alignments is playing a central role when it comes to analyzing high-throughput genomic sequencing data. The GenomicRanges package defines general purpose containers for storing and manipulating genomic intervals and variables defined along a genome. More specialized containers for representing and manipulating short alignments against a reference genome, or a matrix-like summarization of an experiment, are defined in the GenomicAlignments and SummarizedExperiment packages respectively. Both packages build on top of the GenomicRanges infrastructure.
GenomicRanges provides a convenient structure for representing genomic data, and has many built-in functions for manipulating them. The GRanges class represents a collection of genomic ranges that each have a single start and end location on the genome. It can be used to store the location of genomic features such as contiguous binding sites, transcripts, and exons.
Genomic Fragments Visualized through GenomicRanges
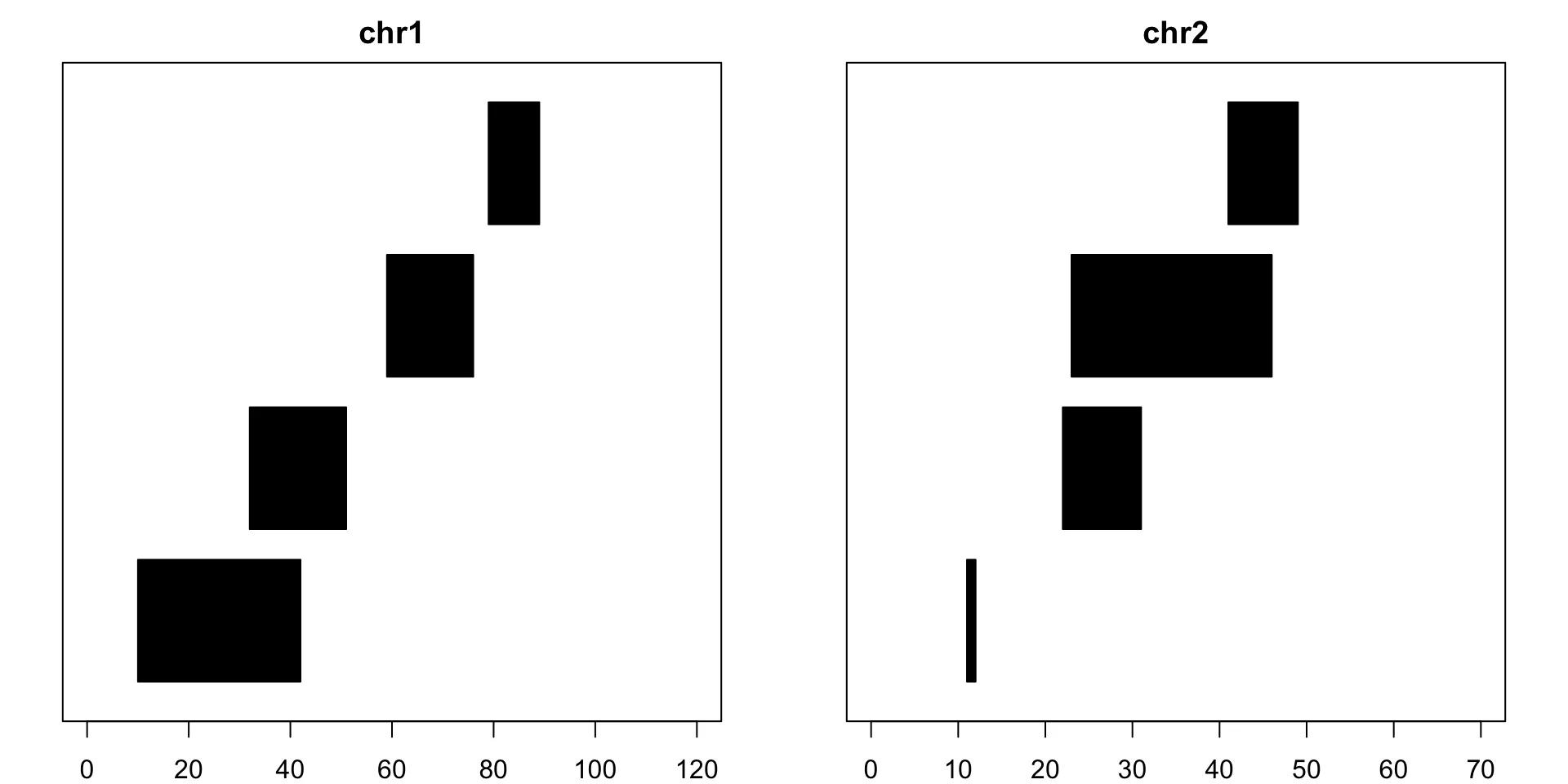
The GPos class is a container for storing a set of genomic positions, that is, genomic ranges of width 1. Even though a GRanges object can be used for that, using a GPos object can be much more memory-efficient, especially when the object contains long runs of adjacent positions.
The GRangesList class is useful for storing genomic features that are inherently compound structures. Whenever genomic features consist of multiple ranges that are grouped by a parent feature, they can be represented as a GRangesList object.
Additional Resources
- Blog: Genomic Ranges: An Introduction to Working with Genomic Data
- Blog: Manipulating Data with dplyr
- Website: Learn More About Domino’s Data Science Platform
- Package Information: Bioconductor Project